GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
Contents:
- Gene and Protein Information
- Previous and Unofficial Names
- Database Links
- Selected 3D Structures
- Enzyme Reaction
- Inhibitors
- Large-scale Ligand Screening Data
- Immunopharmacology Comments
- Immuno Cell Type Associations
- Physiological Consequences of Altering Gene Expression
- Clinically-Relevant Mutations and Pathophysiology
- References
- How to cite this page
Gene and Protein Information  |
||||||
| Species | TM | AA | Chromosomal Location | Gene Symbol | Gene Name | Reference |
| Human | - | 620 | 5q33.3 | ITK | IL2 inducible T cell kinase | |
| Mouse | - | 625 | 11 B1.1 | Itk | IL2 inducible T cell kinase | |
| Rat | - | 626 | 10 q21 | Itk | IL2-inducible T-cell kinase | |
Previous and Unofficial Names  |
| EMT | IL2 inducible T-cell kinase | T-cell-specific kinase | Tcsk | tyrosine-protein kinase ITK/TSK | Tyrosine-protein kinase Lyk |
Database Links  |
|
| Alphafold | Q08881 (Hs), Q03526 (Mm) |
| BRENDA | 2.7.10.2 |
| CATH/Gene3D | 3.30.505.10, 2.30.29.30 |
| ChEMBL Target | CHEMBL2959 (Hs), CHEMBL6067600 (Mm) |
| Ensembl Gene | ENSG00000113263 (Hs), ENSMUSG00000020395 (Mm), ENSRNOG00000006860 (Rn) |
| Entrez Gene | 3702 (Hs), 16428 (Mm), 363577 (Rn) |
| Human Protein Atlas | ENSG00000113263 (Hs) |
| KEGG Enzyme | 2.7.10.2 |
| KEGG Gene | hsa:3702 (Hs), mmu:16428 (Mm), rno:363577 (Rn) |
| OMIM | 186973 (Hs) |
| Orphanet | ORPHA244711 (Hs) |
| Pharos | Q08881 (Hs) |
| RefSeq Nucleotide | NM_005546 (Hs), NM_010583 (Mm), NM_001108825 (Rn) |
| RefSeq Protein | NP_005537 (Hs), NP_034713 (Mm), NP_001102295 (Rn) |
| SynPHARM |
85316 (in complex with BMS-509744) 81635 (in complex with compound 7 [PMID: 22464456]) |
| UniProtKB | Q08881 (Hs), Q03526 (Mm) |
| Wikipedia | ITK (Hs) |
Selected 3D Structures  |
|||||||||||||
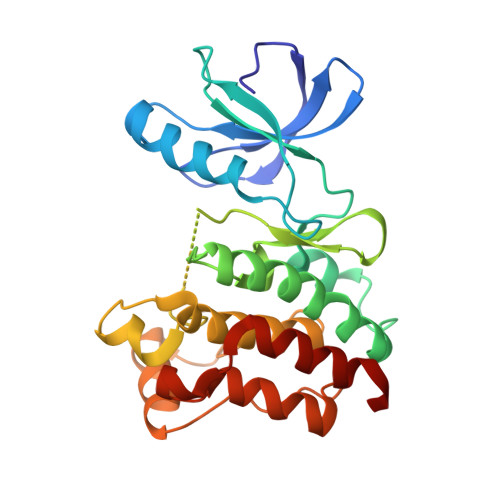
|
|
||||||||||||
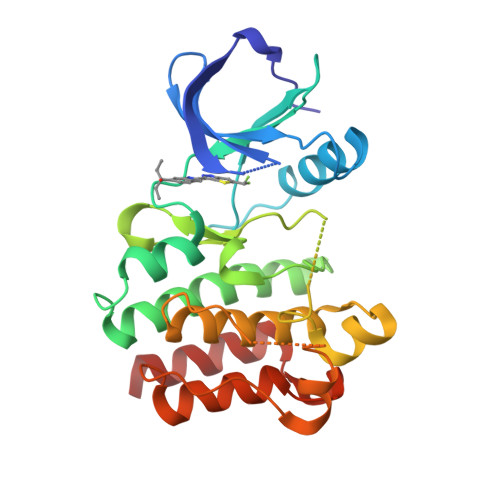
|
|
||||||||||||
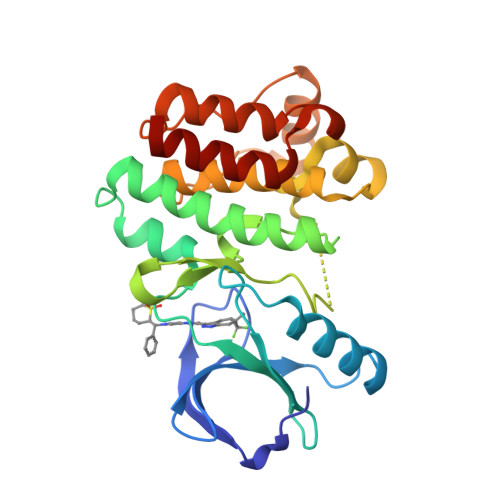
|
|
||||||||||||
Enzyme Reaction  |
||||
|
||||
Download all structure-activity data for this target as a CSV file 
| Inhibitors | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DiscoveRx KINOMEscan® screen  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A screen of 72 inhibitors against 456 human kinases. Quantitative data were derived using DiscoveRx KINOMEscan® platform. http://www.discoverx.com/services/drug-discovery-development-services/kinase-profiling/kinomescan Reference: 7,15 |
 
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Target used in screen: ITK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displaying the top 10 most potent ligands View all ligands in screen » | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
EMD Millipore KinaseProfilerTM screen/Reaction Biology Kinase HotspotSM screen  |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
A screen profiling 158 kinase inhibitors (Calbiochem Protein Kinase Inhibitor Library I and II, catalogue numbers 539744 and 539745) for their inhibitory activity at 1µM and 10µM against 234 human recombinant kinases using the EMD Millipore KinaseProfilerTM service. A screen profiling the inhibitory activity of 178 commercially available kinase inhibitors at 0.5µM against a panel of 300 recombinant protein kinases using the Reaction Biology Corporation Kinase HotspotSM platform. http://www.millipore.com/techpublications/tech1/pf3036 http://www.reactionbiology.com/webapps/main/pages/kinase.aspx Reference: 1,8 |
 
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Target used in screen: Itk/ITK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displaying the top 10 most potent ligands View all ligands in screen » | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Immunopharmacology Comments |
| The TEC family protein tyrosine kinases have been identified as key components of T-cell-receptor activation and signalling. TEC family kinases are expressed predominantly by haematopoietic cells. T cells express ITK, TXK and TEC [2]. ITK inhibition has been extensively expolred as therapeutic option for Th2-driven inflammatory diseases. Despite much effort having been made to discover novel ITK inhibitors, and favourable preclinical discoveries, the majority of the lead compounds have not translated or progressed to the clinic. JTE-051 (structure not disclosed) is currently the only ITK inhibitor under clinical evaluation, and this is being examined in Phase 2 studies as a treatment for rheumatoid arthritis (NCT02919475) and psoriasis (NCT03358290). |
| Cell Type Associations | ||||
|
Physiological Consequences of Altering Gene Expression 
|
||||||||||
|
Clinically-Relevant Mutations and Pathophysiology 
|
||||||||||||||
|
||||||||||||||
References
1. Anastassiadis T, Deacon SW, Devarajan K, Ma H, Peterson JR. (2011) Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat Biotechnol, 29 (11): 1039-45. [PMID:22037377]
2. Berg LJ, Finkelstein LD, Lucas JA, Schwartzberg PL. (2005) Tec family kinases in T lymphocyte development and function. Annu Rev Immunol, 23: 549-600. [PMID:15771581]
3. Brown K, Long JM, Vial SC, Dedi N, Dunster NJ, Renwick SB, Tanner AJ, Frantz JD, Fleming MA, Cheetham GM. (2004) Crystal structures of interleukin-2 tyrosine kinase and their implications for the design of selective inhibitors. J Biol Chem, 279 (18): 18727-32. [PMID:14766749]
4. Burch JD, Barrett K, Chen Y, DeVoss J, Eigenbrot C, Goldsmith R, Ismaili MH, Lau K, Lin Z, Ortwine DF et al.. (2015) Tetrahydroindazoles as Interleukin-2 Inducible T-Cell Kinase Inhibitors. Part II. Second-Generation Analogues with Enhanced Potency, Selectivity, and Pharmacodynamic Modulation in Vivo. J Med Chem, 58 (9): 3806-16. [PMID:25844760]
5. Byrd JC, Harrington B, O'Brien S, Jones JA, Schuh A, Devereux S, Chaves J, Wierda WG, Awan FT, Brown JR et al.. (2016) Acalabrutinib (ACP-196) in Relapsed Chronic Lymphocytic Leukemia. N Engl J Med, 374 (4): 323-32. [PMID:26641137]
6. Das J, Furch JA, Liu C, Moquin RV, Lin J, Spergel SH, McIntyre KW, Shuster DJ, O'Day KD, Penhallow B et al.. (2006) Discovery and SAR of 2-amino-5-(thioaryl)thiazoles as potent and selective Itk inhibitors. Bioorg Med Chem Lett, 16 (14): 3706-12. [PMID:16682193]
7. Davis MI, Hunt JP, Herrgard S, Ciceri P, Wodicka LM, Pallares G, Hocker M, Treiber DK, Zarrinkar PP. (2011) Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol, 29 (11): 1046-51. [PMID:22037378]
8. Gao Y, Davies SP, Augustin M, Woodward A, Patel UA, Kovelman R, Harvey KJ. (2013) A broad activity screen in support of a chemogenomic map for kinase signalling research and drug discovery. Biochem J, 451 (2): 313-28. [PMID:23398362]
9. Hain A, Krämer M, Linka RM, Nakhaei-Rad S, Ahmadian MR, Häussinger D, Borkhardt A, Münk C. (2018) IL-2 Inducible Kinase ITK is Critical for HIV-1 Infection of Jurkat T-cells. Sci Rep, 8 (1): 3217. [PMID:29453458]
10. Herdemann M, Weber A, Jonveaux J, Schwoebel F, Stoeck M, Heit I. (2011) Optimisation of ITK inhibitors through successive iterative design cycles. Bioorg Med Chem Lett, 21 (6): 1852-6. [PMID:21316219]
11. Hodson R, Beausoleil AM. (2018) Compounds and methods for modulating interleukin-2-inducible t-cell kinase. Patent number: WO2018089261A2. Assignee: Corvus Pharmaceuticals, Inc.. Priority date: 03/11/2017. Publication date: 17/05/2018.
12. Jacobsen EJ, Anderson DR, Blinn JR, Hockerman SL, Heier R, Mukherjee P. (2025) Pyrrolopyrimidine ITK inhibitors. Patent number: WO2020033955A1. Assignee: Aclaris Therapeutics, Inc.. Priority date: 24/10/2023. Publication date: 13/05/2025.
13. Kaul A, Hope H, Xu C, Basavalingappa R, Binz SK, Boily C, Bradley Z, Burt D, Emanuel C, Fairchild J et al.. (2025) Characterization of the dual ITK/JAK3 small molecule covalent inhibitor ATI-2138. J Pharmacol Exp Ther, 392 (2): 100054. [PMID:40023606]
14. McLean LR, Zhang Y, Zaidi N, Bi X, Wang R, Dharanipragada R, Jurcak JG, Gillespy TA, Zhao Z, Musick KY et al.. (2012) X-ray crystallographic structure-based design of selective thienopyrazole inhibitors for interleukin-2-inducible tyrosine kinase. Bioorg Med Chem Lett, 22 (9): 3296-300. [PMID:22464456]
15. Wodicka LM, Ciceri P, Davis MI, Hunt JP, Floyd M, Salerno S, Hua XH, Ford JM, Armstrong RC, Zarrinkar PP et al.. (2010) Activation state-dependent binding of small molecule kinase inhibitors: structural insights from biochemistry. Chem Biol, 17 (11): 1241-9. [PMID:21095574]
16. Yao X, Sun X, Jin S, Yang L, Xu H, Rao Y. (2019) Discovery of 4-Aminoquinoline-3-carboxamide Derivatives as Potent Reversible Bruton's Tyrosine Kinase Inhibitors for the Treatment of Rheumatoid Arthritis. J Med Chem, 62 (14): 6561-6574. [PMID:31260299]
17. Zhong Y, Dong S, Strattan E, Ren L, Butchar JP, Thornton K, Mishra A, Porcu P, Bradshaw JM, Bisconte A et al.. (2015) Targeting interleukin-2-inducible T-cell kinase (ITK) and resting lymphocyte kinase (RLK) using a novel covalent inhibitor PRN694. J Biol Chem, 290 (10): 5960-78. [PMID:25593320]
18. Zhou W, Ercan D, Chen L, Yun CH, Li D, Capelletti M, Cortot AB, Chirieac L, Iacob RE, Padera R et al.. (2009) Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature, 462 (7276): 1070-4. [PMID:20033049]
How to cite this page
Tec family: IL2 inducible T cell kinase. Last modified on 29/07/2025. Accessed on 03/05/2026. IUPHAR/BPS Guide to PHARMACOLOGY, https://www.guidetomalariapharmacology.org/GRAC/ObjectDisplayForward?objectId=2046.