GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
|
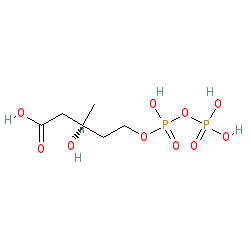
Synonyms: mevalonate 5-diphosphate
Compound class:
Metabolite
Ligand Activity Visualisation ChartsThese are box plot that provide a unique visualisation, summarising all the activity data for a ligand taken from ChEMBL and GtoPdb across multiple targets and species. Click on a plot to see the median, interquartile range, low and high data points. A value of zero indicates that no data are available. A separate chart is created for each target, and where possible the algorithm tries to merge ChEMBL and GtoPdb targets by matching them on name and UniProt accession, for each available species. However, please note that inconsistency in naming of targets may lead to data for the same target being reported across multiple charts. ✖ |
|
|||||||||||||||||||||||||||||||||||
| References |
|
1. Herdendorf TJ, Miziorko HM. (2006)
Phosphomevalonate kinase: functional investigation of the recombinant human enzyme. Biochemistry, 45 (10): 3235-42. [PMID:16519518] |
|
2. Herdendorf TJ, Miziorko HM. (2007)
Functional evaluation of conserved basic residues in human phosphomevalonate kinase. Biochemistry, 46 (42): 11780-8. [PMID:17902708] |
|
3. Michihara A, Sawamura M, Nara Y, Ikeda K, Yamori Y. (1997)
Purification and characterization of two mevalonate pyrophosphate decarboxylases from rat liver: a novel molecular species of 37 kDa. J Biochem, 122 (3): 647-54. [PMID:9348097] |
|
4. Qiu Y, Gao J, Guo F, Qiao Y, Li D. (2007)
Mutation and inhibition studies of mevalonate 5-diphosphate decarboxylase. Bioorg Med Chem Lett, 17 (22): 6164-8. [PMID:17888661] |
|
5. Qiu Y, Li D. (2006)
Inhibition of mevalonate 5-diphosphate decarboxylase by fluorinated substrate analogs. Biochim Biophys Acta, 1760 (7): 1080-7. [PMID:16626865] |
|
6. Toth MJ, Huwyler L, Park J. (1996)
Purification of rat liver mevalonate pyrophosphate decarboxylase. Prep Biochem Biotechnol, 26 (1): 47-51. [PMID:8744421] |
|
7. Voynova NE, Fu Z, Battaile KP, Herdendorf TJ, Kim JJ, Miziorko HM. (2008)
Human mevalonate diphosphate decarboxylase: characterization, investigation of the mevalonate diphosphate binding site, and crystal structure. Arch Biochem Biophys, 480 (1): 58-67. [PMID:18823933] |