
GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
Calcium- and sodium-activated potassium channels (KCa, KNa): Introduction
Adapted from the original 2005 article in Pharmacological Reviews [17], and update published in 2017 [7].
The second major group of six/seven transmembrane potassium-selective channels consists of the KCa channels (for reviews, see [2,9-11,16]), and Table 1 shows the International Union of Pharmacology (IUPHAR) names of the members of this group together with their HUGO Gene Nomenclature Committee (HGNC) designations and other commonly used names. The phylogenetic trees in Fig. 1 illustrate the fact that these channels form two well defined but only distantly related groups.
TABLE 1 - KCa channels: IUPHAR names of the members of the KCa group of potassium channels are shown, together with their HGNC designations and other commonly used names. BK, big-conductance K+ channel; SK, small-conductance K+ channel; IK, intermediate-conductance K+ channel.
| ||||||||||||||||||||||||||||||
One of these groups (Fig. 1A) includes the three "small-conductance" KCa channels (KCa2.1, 2.2, and 2.3) [8] and the "intermediate-conductance" channel KCa3.1 [5-6]. These channels are voltage-insensitive and are activated by low concentrations of internal Ca2+ (<1.0 μM), in contrast to KCa1.1 (KCNMA1, Slo1), which is activated by both voltage and internal Ca2+. The three small-conductance KCa channels are sensitive to block by apamin (100 pM-10 nM), which distinguishes them from all other KCa channels. Both small- and intermediate-conductance KCa channels play important roles in many processes involving Ca2+-dependent signaling in both electrically excitable and nonexcitable cells. They do not bind Ca2+ directly but rather detect Ca2+ by virtue of calmodulin, which is constitutively bound to the C-terminal region [4,19]. Binding of calcium to this calmodulin results in conformational changes that are in turn responsible for channel gating [14].
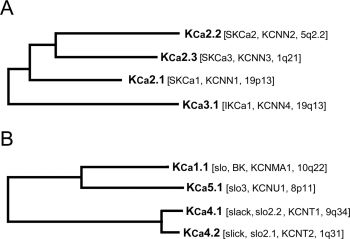
FIG. 1. Phylogenetic tree for KCa channels. A, KCa2/3 group. B, KCa1/4/5 group. Amino acid sequence alignments for the known members of the human KCa family were created using CLUSTALW. Analysis by maximum parsimony using PAUP* resulted in the above tree. International Union of Pharmacology and HUGO Gene Nomenclature Committee names of the genes are shown together with their chromosomal location.
The tree shown in Fig. 1B illustrates the sequence relationships within the second group of KCa channels, which includes KCa1.1 (Slo or Slo1), KCa4.1 (Slack or Slo2.2), KCa4.2 (Slick or Slo2.1), and KCa5.1 (Slo3). KCa1.1 has been extensively studied in the brain, cochlea, and muscle, and alternate splicing of its mRNA is known to produce considerable functional diversity [3,18]. Unlike the KCa2 and KCa3 channels, binding of calcium by KCa1.1 is not dependent on its association with calmodulin but is thought to be mediated by at least three divalent cation binding sites in the cytoplasmic carboxyl domain of each channel subunit. Two independent high-affinity Ca2+ binding sites are formed by a negatively charged segment in the distal carboxyl terminal portion, termed the "calcium bowl" [12] and within the first RCK domain encoded by the proximal C-terminal portion [1,20]. A third low-affinity divalent cation binding site is also found in the first RCK domain [15], which contributes to activation by Mg2+ and Ca2+ at high concentrations (>1 mM).
The three other members of this group, KCa4.1, 4.2, and 5.2 [6,13,21], were all included in the KCa nomenclature since they all are clearly members of this structurally related group of genes. However, much more is now known about the functional properties of the members of this gene family than was known when these names were assigned several years ago, and this presents a possible conundrum for a nomenclature based on functional rather than structural similarity. Unlike the founding member KCa1.1, which is in fact activated by internal Ca2+, none of the other members of this group seems to be similarly Ca2+-activated. In fact, for the most part, these three are insensitive to internal Ca2+. KCa4.2 and KCa4.1 are activated by internal Na+ and Cl- [21], and KCa5.1 is activated by internal alkalization (OH-) [13]. Therefore, although they are structurally related to KCa1.1, these three channels cannot correctly be described as "calcium-activated" channels based on functional criteria. This may be a subject for discussion among researchers in this field and those bodies responsible for standardizing gene nomenclature.
References
1. Bao L, Rapin AM, Holmstrand EC, Cox DH. (2002) Elimination of the BK(Ca) channel's high-affinity Ca(2+) sensitivity. J Gen Physiol, 120 (2): 173-89. [PMID:12149279]
2. Cox DH. (2005) The BKCa channel's Ca2+-binding sites, multiple sites, multiple ions. J Gen Physiol, 125 (3): 253-5. [PMID:15738047]
3. Faber ES, Sah P. (2003) Calcium-activated potassium channels: multiple contributions to neuronal function. Neuroscientist, 9 (3): 181-94. [PMID:15065814]
4. Fanger CM, Ghanshani S, Logsdon NJ, Rauer H, Kalman K, Zhou J, Beckingham K, Chandy KG, Cahalan MD, Aiyar J. (1999) Calmodulin mediates calcium-dependent activation of the intermediate conductance KCa channel, IKCa1. J Biol Chem, 274 (9): 5746-54. [PMID:10026195]
5. Ishii TM, Silvia C, Hirschberg B, Bond CT, Adelman JP, Maylie J. (1997) A human intermediate conductance calcium-activated potassium channel. Proc Natl Acad Sci USA, 94 (21): 11651-6. [PMID:9326665]
6. Joiner WJ, Wang LY, Tang MD, Kaczmarek LK. (1997) hSK4, a member of a novel subfamily of calcium-activated potassium channels. Proc Natl Acad Sci USA, 94 (20): 11013-8. [PMID:9380751]
7. Kaczmarek LK, Aldrich RW, Chandy KG, Grissmer S, Wei AD, Wulff H. (2017) International Union of Basic and Clinical Pharmacology. C. Nomenclature and Properties of Calcium-Activated and Sodium-Activated Potassium Channels. Pharmacol Rev, 69 (1): 1-11. [PMID:28267675]
8. Kohler M, Hirschberg B, Bond CT, Kinzie JM, Marrion NV, Maylie J, Adelman JP. (1996) Small-conductance, calcium-activated potassium channels from mammalian brain. Science, 273 (5282): 1709-14. [PMID:8781233]
9. Lingle CJ. (2002) Setting the stage for molecular dissection of the regulatory components of BK channels. J Gen Physiol, 120 (3): 261-5. [PMID:12198086]
10. Magleby KL. (2003) Gating mechanism of BK (Slo1) channels: so near, yet so far. J Gen Physiol, 121 (2): 81-96. [PMID:12566537]
11. Moczydlowski EG. (2004) BK channel news: full coverage on the calcium bowl. J Gen Physiol, 123 (5): 471-3. [PMID:15111642]
12. Schreiber M, Salkoff L. (1997) A novel calcium-sensing domain in the BK channel. Biophys J, 73 (3): 1355-63. [PMID:9284303]
13. Schreiber M, Wei A, Yuan A, Gaut J, Saito M, Salkoff L. (1998) Slo3, a novel pH-sensitive K+ channel from mammalian spermatocytes. J Biol Chem, 273 (6): 3509-16. [PMID:9452476]
14. Schumacher MA, Rivard AF, Bächinger HP, Adelman JP. (2001) Structure of the gating domain of a Ca2+-activated K+ channel complexed with Ca2+/calmodulin. Nature, 410 (6832): 1120-4. [PMID:11323678]
15. Shi J, Krishnamoorthy G, Yang Y, Hu L, Chaturvedi N, Harilal D, Qin J, Cui J. (2002) Mechanism of magnesium activation of calcium-activated potassium channels. Nature, 418 (6900): 876-80. [PMID:12192410]
16. Stocker M. (2004) Ca(2+)-activated K+ channels: molecular determinants and function of the SK family. Nat Rev Neurosci, 5 (10): 758-70. [PMID:15378036]
17. Wei AD, Gutman GA, Aldrich R, Chandy KG, Grissmer S, Wulff H. (2005) International Union of Pharmacology. LII. Nomenclature and molecular relationships of calcium-activated potassium channels. Pharmacol Rev, 57 (4): 463-72. [PMID:16382103]
18. Weiger TM, Hermann A, Levitan IB. (2002) Modulation of calcium-activated potassium channels. J Comp Physiol A Neuroethol Sens Neural Behav Physiol, 188 (2): 79-87. [PMID:11919690]
19. Xia XM, Fakler B, Rivard A, Wayman G, Johnson-Pais T, Keen JE, Ishii T, Hirschberg B, Bond CT, Lutsenko S et al.. (1998) Mechanism of calcium gating in small-conductance calcium-activated potassium channels. Nature, 395 (6701): 503-7. [PMID:9774106]
20. Xia XM, Zeng X, Lingle CJ. (2002) Multiple regulatory sites in large-conductance calcium-activated potassium channels. Nature, 418 (6900): 880-4. [PMID:12192411]
21. Yuan A, Santi CM, Wei A, Wang ZW, Pollak K, Nonet M, Kaczmarek L, Crowder CM, Salkoff L. (2003) The sodium-activated potassium channel is encoded by a member of the Slo gene family. Neuron, 37 (5): 765-73. [PMID:12628167]







